Our faculty in the Agricultural and Biological Sciences Department pursue excellence in scholarship—guiding, mentoring and challenging their students to, likewise, explore the questions that intrigue them. Here you can read about the wide range of Agricultural and Biological Sciences faculty research interests; you can also access some recent publications to get a better understanding of the type of research our faculty are doing.
Dr. Brett Berke, PhD
University of Iowa
bberke@truman.edu
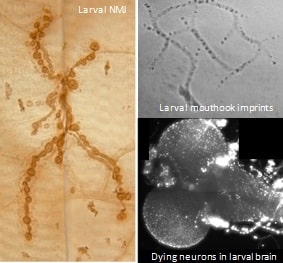
Research in our lab uses the genetic tool kit of fruit flies (Drosophila melanogaster) to identify molecular mechanisms of synapse formation, animal behavior, and neurodegeneration. We study how mutations alter development of the larval neuromuscular junction and how they affect distinct aspects of larval crawling behavior. We also use mutations to address how neurons die in fly models of neurological illness.
Check out these papers to learn more about research in the Berke lab:
A. Galbraith, S. Leone, K. Stuart, J. Emery, M. K. Renkemeyer, N. Pritchett, L. Galbraith, H. Stuckmeyer, and B. Berke. 2021. Reducing the expression of the Numb adaptor protein in neurons increases the searching behavior of Drosophila larvae. microPublication Biology.
M. Thies and B. Berke. 2020. A role for the Fem-1 gene of Drosophila melanogaster in adult courtship. bioRxiv.
E. M. McNeill, C. Thompson, B. Berke, V. T. Chou, J. Rusch, A. Duckworth, J. DeProto, A. Taylor, J. Gates, F. Gertler, H. Keshishian, and D. Van Vactor. 2020. Drosophila enabled promotes synapse morphogenesis and regulates active zone form and function. Neural Development 15: 1-13.
B. Berke, L. Le, and H. Keshishian. 2019. Target-Dependent Retrograde Signaling Mediates Synaptic Plasticity at the Drosophila Neuromuscular Junction. Developmental Neurobiology 79: 895-912.
B. Berke, J. Wittnam, and H. Keshishian. 2013. Retrograde BMP Sign.aling at the Synapse: A Permissive Signal for Synapse Maturation and Activity-Dependent Plasticity. Journal of Neuroscience 33: 17937-17950.
My research interests include the scholarship of teaching and learning, science education, and teacher professional development strategies. I also study science anxiety, motivation and attitude.
Check out these papers to learn more about research in the Berke lab:
Udo M.K., G.P. Ramsey, J.V. Mallow. 2004. Science Anxiety and Gender in Students Taking General Education Science Courses. Journal of Science Education and Technology 13: 435-446.
Mallow, J.V. 2006. Science anxiety: research and action. Handbook of college science teaching, pp.3-14.
Dr. Maria Beatriz de Souza Cortez
University of Florida (Ph.D)
University of Campinas (Masters)
mbscortez@truman.edu
My interest in conservation mostly relates to a relatively new field named biocultural conservation, which aims to align the protection of flora and fauna with the needs and culture of locals and indigenous populations.
Most of my research has been focused on Brazilian plants, more specifically those occurring in the campos rupestres. The grassy and rocky landscape of this vegetation type is ripe with biodiversity. Amongst the most important plants in the region are the sempre vivas, a group of plants that have been harvested by local communities as a means of subsistence.
Finally, I am also interested in expanding outreach efforts that connect university and community.
Check out these papers to learn more about research in the Beatriz lab:
Soltis, D.E., Smocovitis, V.B., Pham, K.K., Cortez, M.B.S., Smith, A.L. and Soltis, P.S. (2023) Rethinking the Ph.D. dissertation in botany: Widening the circle. American Journal of Botany, 110: e16136.
De Souza Cortez, M.B.,Folk, R.A., Grady, C.J., Spoelhof, J.P., Smith, S.A., Soltis, D.E., Soltis, P.S., (2021) Is the age of plant communities predicted by the age, stability and soil composition of the underlying landscapes? An investigation of OCBILs, Biological Journal of the Linnean Society, 133(2): 297–316.
Cortez, M.B.S., Sforça, D.A., Alves, F.M., Vidal, J.D., Alves-Pereira, A., Mori, G.M., Andreotti, I.A., Do Nascimento, J.E., Bittrich, V., Zucchi, M.I., Amaral, M.C.E., Souza, A.P., (2019) Elucidating the Clusia criuva species ‘complex’: cryptic taxa can exhibit great genetic and geographical variation, Botanical Journal of the Linnean Society, 190(1): 67–82.
Dr. Laura Fielden, PhD
University of Natal, Pietermaritzberg, South Africa
lfielden@truman.edu

Check out these papers to learn more about research in the Fielden lab:
Khokhlova I.S., L.J. Fielden, A. Degen, and B.R. Krasnov. 2012. Ectoparasite fitness in auxiliary hosts: Phylogenetic distance from a principle host matters. Journal of Evolutionary Biology 25: 2005-2013.
Khokhlova I.S., L.J. Fielden, J.B. Williams, A. Degen, and B.R. Krasnov. 2013. Energy expenditure for egg production in arthropod ectoparasites: The effect of host species. Parasitology 140: 1070-1077.
Khokhlova I.S., S. Pilosof, L.J. Fielden, A. Degen, and B.R. Krasnov. 2014. A trade-off between quantity and quality of offspring in haematophagous ectoparasites: the effect of level of specialization. Journal of Animal Ecology 83: 397-405.
Dr. Stephanie Foré, PhD
North Carolina State University of Miami
sfore@truman.edu

Check out these papers to learn more about research in the Foré lab:
Fore, S.A., M.F. Mangan, A.M. Mantia, J.T. Kolok and H.J. Kim. 2023. Multiple physiological cohorts comprise seasonal activity of wild Amblyomma americanum (Acri: Ixodidae) nymphs. Ticks and Tick-borne Diseases 14:102091 Fore, S.A. and H.J. Kim. 2021. Pattern of early questing in unfed larval Amblyomma americanum from twelve years of monitoring in northern Missouri, USA. International Journal of Acarology DOI: 10.1080/01647954.2021.1966501
Mangan, M., S.A. Fore and H.J. Kim. 2020. Using haem concentration as a metric of physiological age to infer demographic structure in natural field and forest populations of host-seeking Amblyomma americanum adults.International Journal of Acarology, 46:258-262.
DOI: 10.1080/01647954.2020.1758776
Kaizer A.M., S.A. Foré , H.-J. Kim, and E.C. York. 2015. Modeling the biotic and abiotic factors that describe the number of active off-host Amblyomma americanum larvae. Journal of Vector Ecology 40: 1-10.
Bouzek, D.C., S.A. Foré, J.G. Bevell, and H.-J. Kim. 2013. A conceptual model of the Amblyomma americanum life cycle in northeast Missouri. Journal of Vector Ecology 38: 74-81.
Dallas, T.A., S.A. Foré, and H.-J. Kim. 2012. Modeling the influence of Peromyscus leucopus body mass, sex and habitat on immature Dermacentor variabilis burden. Journal of Vector Ecology 37: 338-341.
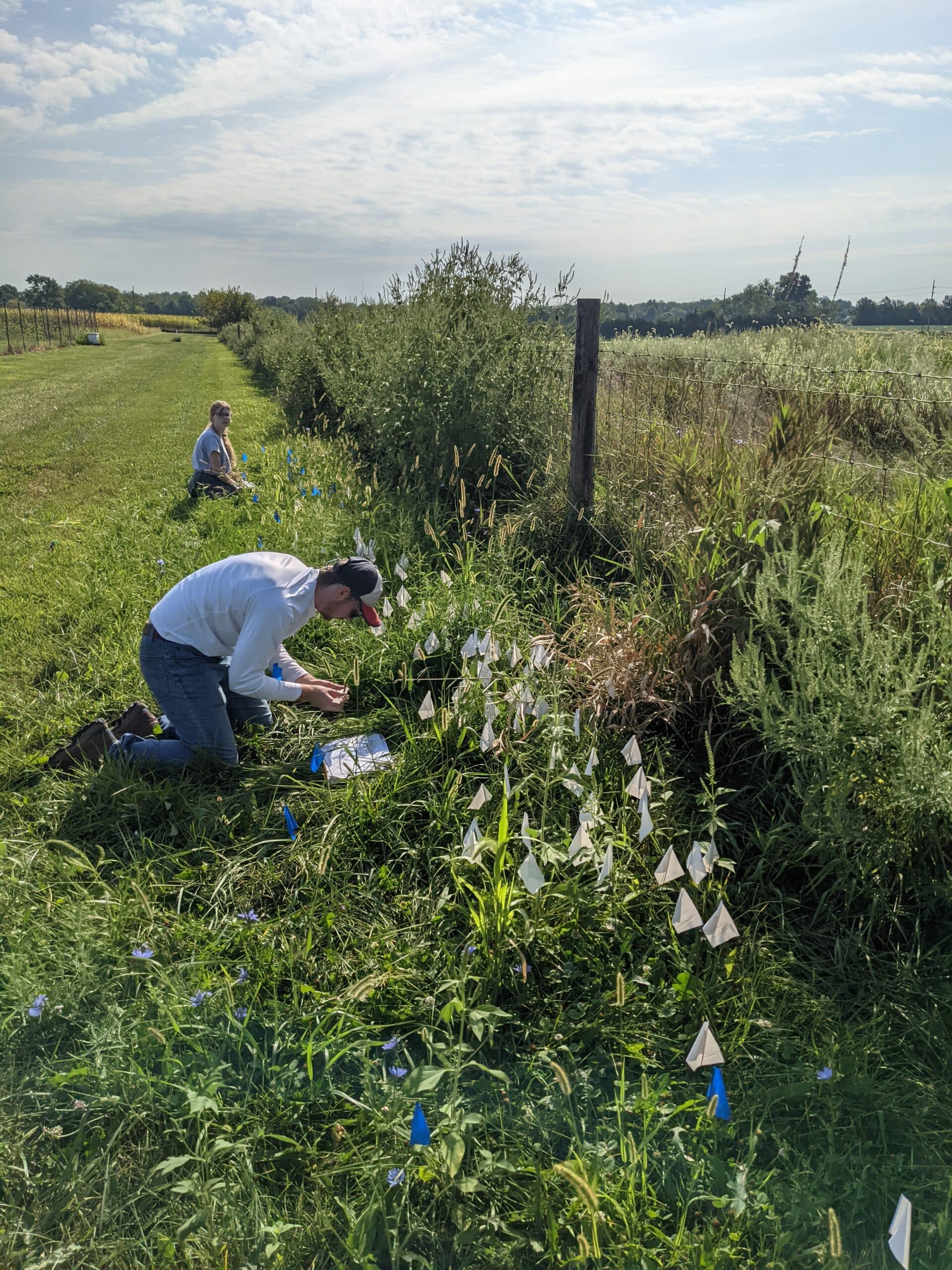
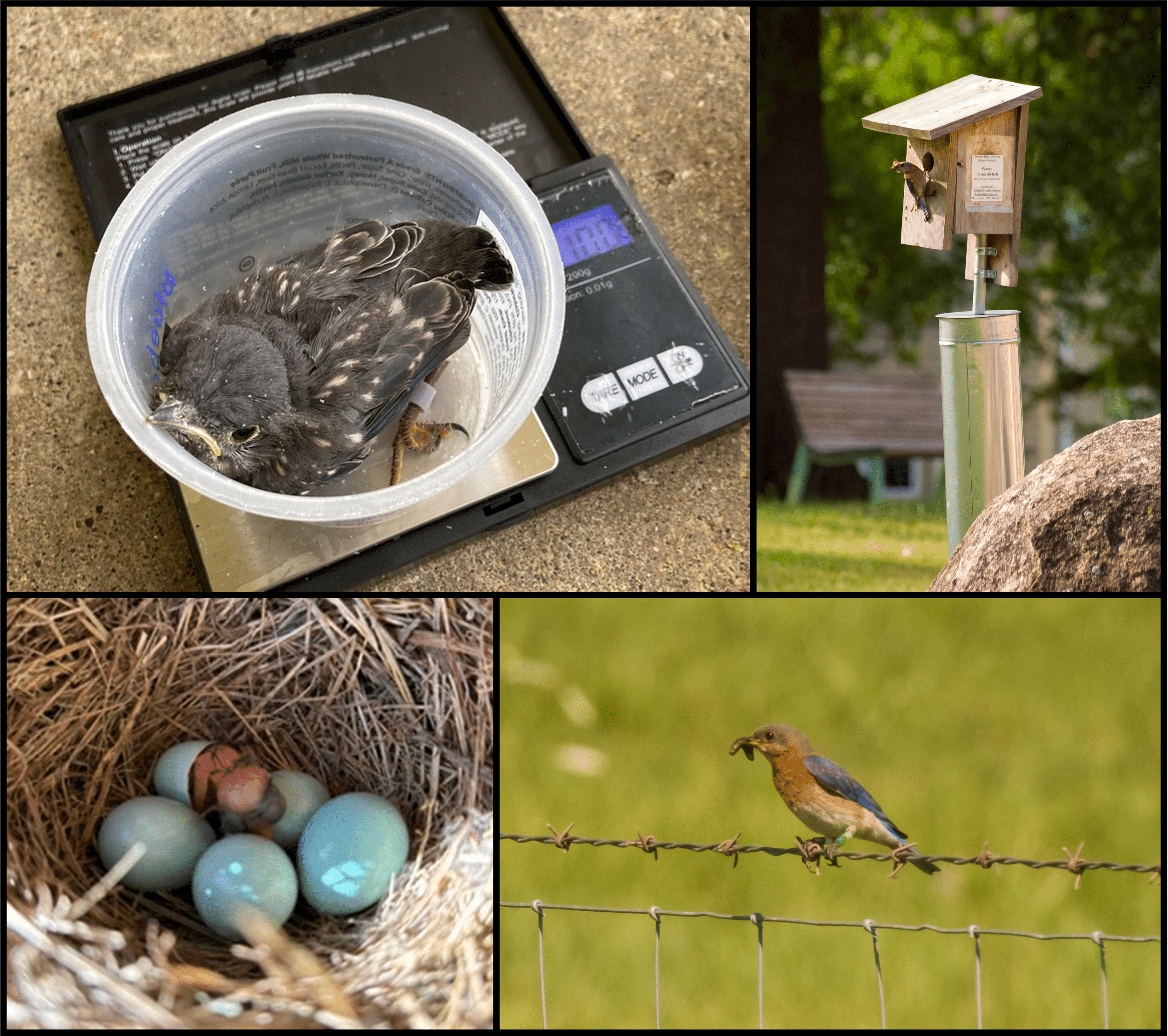
Student-led projects in the Hubbard Lab have taken many forms and students are encouraged to explore their interests within the areas of ornithology, behavior, ecology, and evolution. For example, students have examined plumage color measurement methodology, responses and habituation to real and mimicked predator calls at bird feeders, and prevalence of window strikes on Truman’s campus.
Check out these papers to learn more about research in the Hubbard lab:
Hubbard, J.K. & Z.W.D. Williard. 2023. Spectra of feather samples are impacted by the substrate color against which they are measured. The Wilson Journal of Ornithology 135: 1-9.
Hubbard, J.K., J.A.C. Uy, M.E. Hauber, H.E. Hoekstra, R.J. Safran. 2012 Vertebrate pigmentation from underlying genes to adaptive function. Trends in Genetics 26: 231-239.
Hubbard J.K. B.R. Jenkins, R.J. Safran. 2015. Quantitative genetics of plumage color: Lifetime effects of early nest environment on a colorful sexual signal. Ecology and Evolution 5: 3436-3449.
Students working in my lab use PCR-based data collection techniques and modern statistical methods to address questions about natural populations. Examples include describing contemporary genetics structure in natural populations using DNA fingerprints and presence/absence of disease-causing fungi in local populations.
Check out these papers to learn more about research in the Hudman lab:
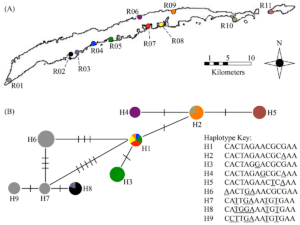
Pasachnik, S.A. and S.P. Hudman. 2016. Conservation genetics of Roatán spiny-tailed iguanas, Ctenosaura oedirhina. Herpetological Conservation and Biology, 11: 187-196.
Lennon C., S.P. Hudman, C.E. Montgomery. 2014. Assessment of Batrachochytrium dendrobatidis infection level in amphibians of Wakonda State Park, Missouri, USA. Herpetological Review, 45: 40.
Hudman S.P. and K.B. Gido. 2013. Multi-scale effects of impoundments on genetic structure of creek chub (Semotilus atromaculatus) in the Kansas River basin. Freshwater Biology 58: 441-453.
Dr. Ali Hussein
Oklahoma State University
ahussein@truman.edu
Check out these papers to learn more about research in the Hussein lab:
Hussein, A. H., R. Puchala, I. Portugal, T. A. Gipson, B. K. Wilson, and A. L. Goetsch. 2020. Effects of water restriction on feed intake, digestion, and energy utilization by mature
female St. Croix sheep. Vet. Anim. Sci. 10:1-6. doi:10.1016/j.vas.2020.100132
Hussein, A. H., E. D. Batista, M. D. Miesner, and E. C. Titgemeyer. 2016. Effect of ruminal
ammonia supply on lysine utilization by growing steers. J. Anim. Sci. 94:656–664.
doi:10.2527/jas.2015-9717
Dr. Bob Johnson
University of California, Davis
bjohnson@truman.edu

Check out these papers to learn more about research in the Hussein lab:
B. Johnson, F. Niederholzer, L. Milliron and D.M. Rizzo, Improved understanding of the threat posed by Phellinus heart-rot in California prune orchards, Acta Hortic. 1322. ISHS 2021, Proc. XII Int. Symposium on Plum and Prune Genetics, Breeding and Pomology.
D.A. Hernandeza, B. Johnson and D.M. Rizzo, Management and biocontrol of Phellinus heart-rot in California prunes, Acta Hortic. 1322. ISHS 2021, Proc. XII Int. Symposium on Plum and Prune Genetics, Breeding and Pomology.
Dr. Josh Kraft
Purdue University
jkraft@truman.edu

Check out this paper to learn more about research in the Ma lab:
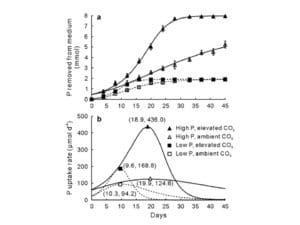
Zhong, M.A., J. Flynn, G. Libra, Z. Shi. 2018. Elevated CO2 accelerates depletion of phosphorus by common bean (Phaseolus vulgaris) in association with altered leaf biochemical properties. Pedosphere 28: 422-429.
There have also been student-led projects in the lab, such as the role of specific genes in the activation of C. elegans sperm or the role of a heme transporter in the adult epidermis. For students who wish to develop their own project, I work closely with them to design a methodology that is feasible.
Check out these papers to learn more about research in the Maiden lab:
Prins, R., Windsor, P., Miller III, B. R., & Maiden, S. (2022). Alleles of unc‐33/CRMP exhibit defects during Caenorhabditis elegans epidermal morphogenesis. Developmental Dynamics, 251(10), 1741-1753.
Wieberg, S., Euwer, H., Gerst, A., & Maiden, S. L. (2021). Synergistic effects of hmp-2/β-catenin and sma-1/βH-spectrin on epidermal morphogenesis in Caenorhabditis elegans. microPublication Biology, 2021.
Garzanelli, J., & Maiden, S. (2020). Exposing a novel genetic interaction between unc-33/CRMP and hmp-2/β-catenin during Caenorhabditis elegans embryogenesis. microPublication Biology, 2020.
Maiden, S.L., N. Harrison, J. Keegan, B. Cain, A.M. Lynch, J. Pettitt, and J. Hardin. 2013. Specific conserved C-terminal amino acids of Caenorhabditis elegans HMP-1/α-catenin modulate F-actin binding independently of vinculin. Journal of Biological Chemistry 288: 5694-5706.
Maiden, S.L., Y.I. Petrova, and B.M. Gumbiner. 2016. Microtubules inhibit E-cadherin adhesive activity by maintaining phosphorylated p120-catenin in a colon carcinoma cell model. PloS one 11: e0148574.
Dr. Hajeewaka Mendis
Florida State University
hmendis@truman.edu
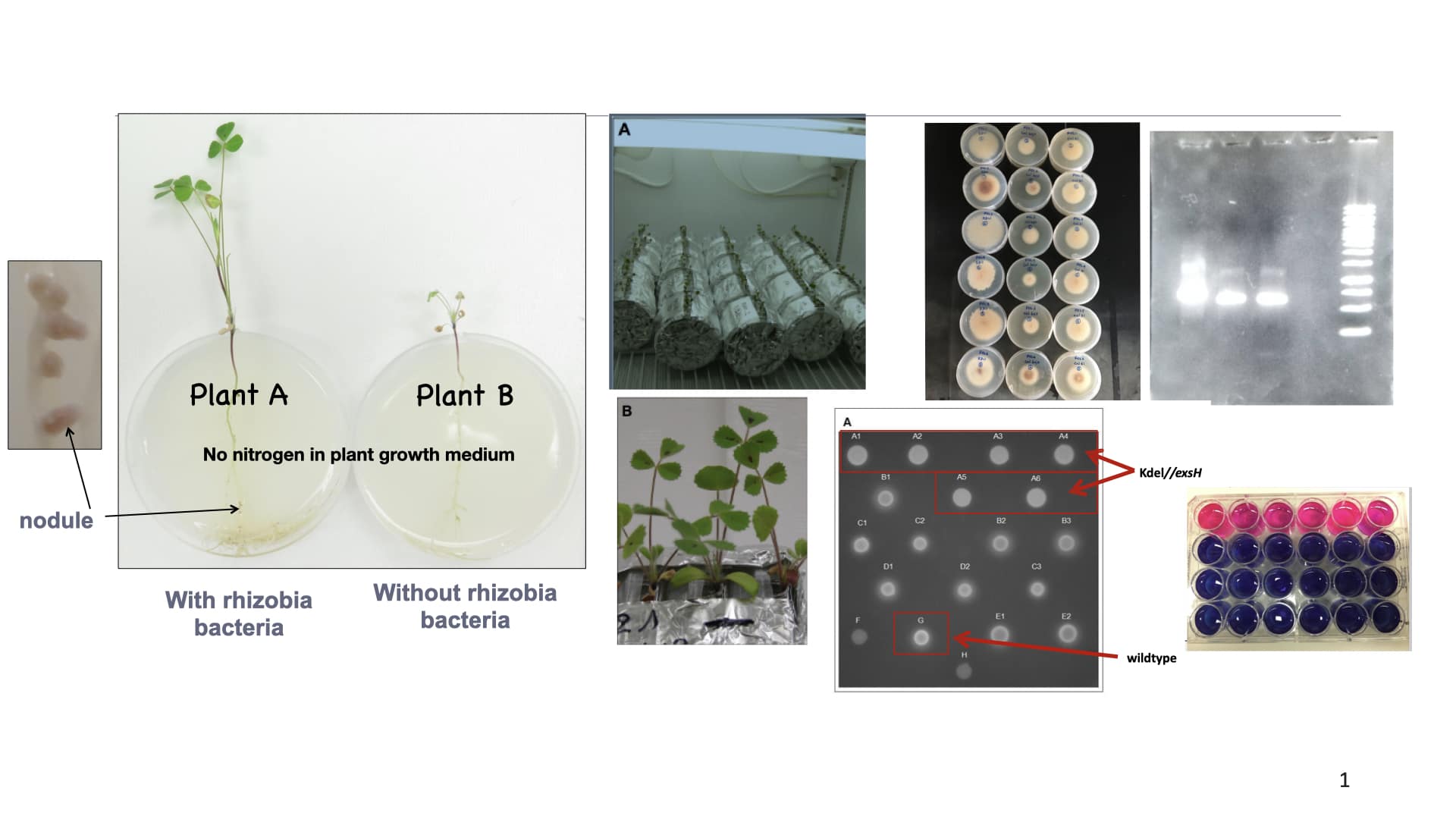

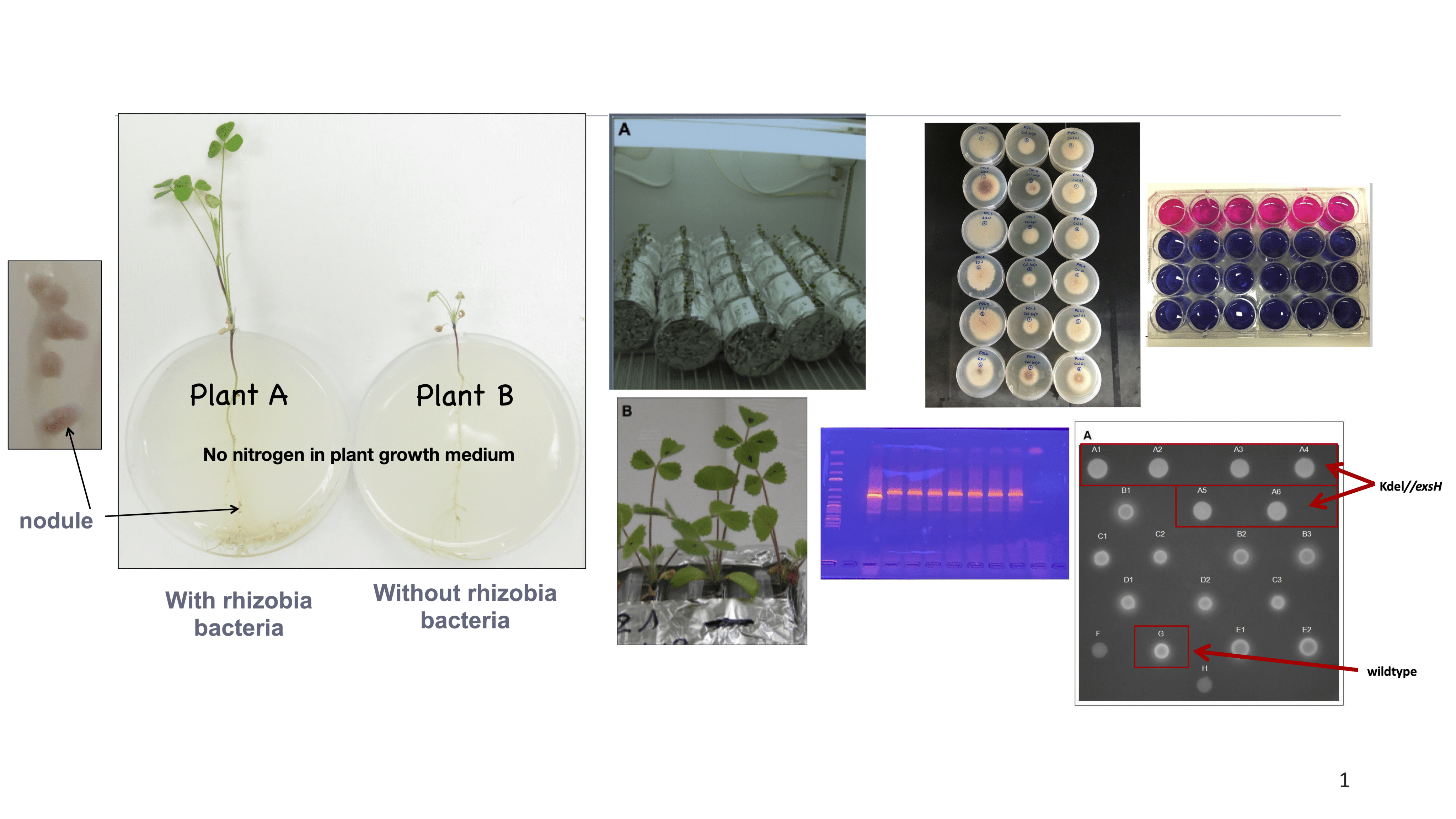
- The function of bacterial exopolysaccharides during root infection and symbiosis of nitrogen fixing bacteria Sinorhizobium meliloti and legume plant Medicago truncatula.
- Importance of bacterial exopolysaccharides in biofilm formation.
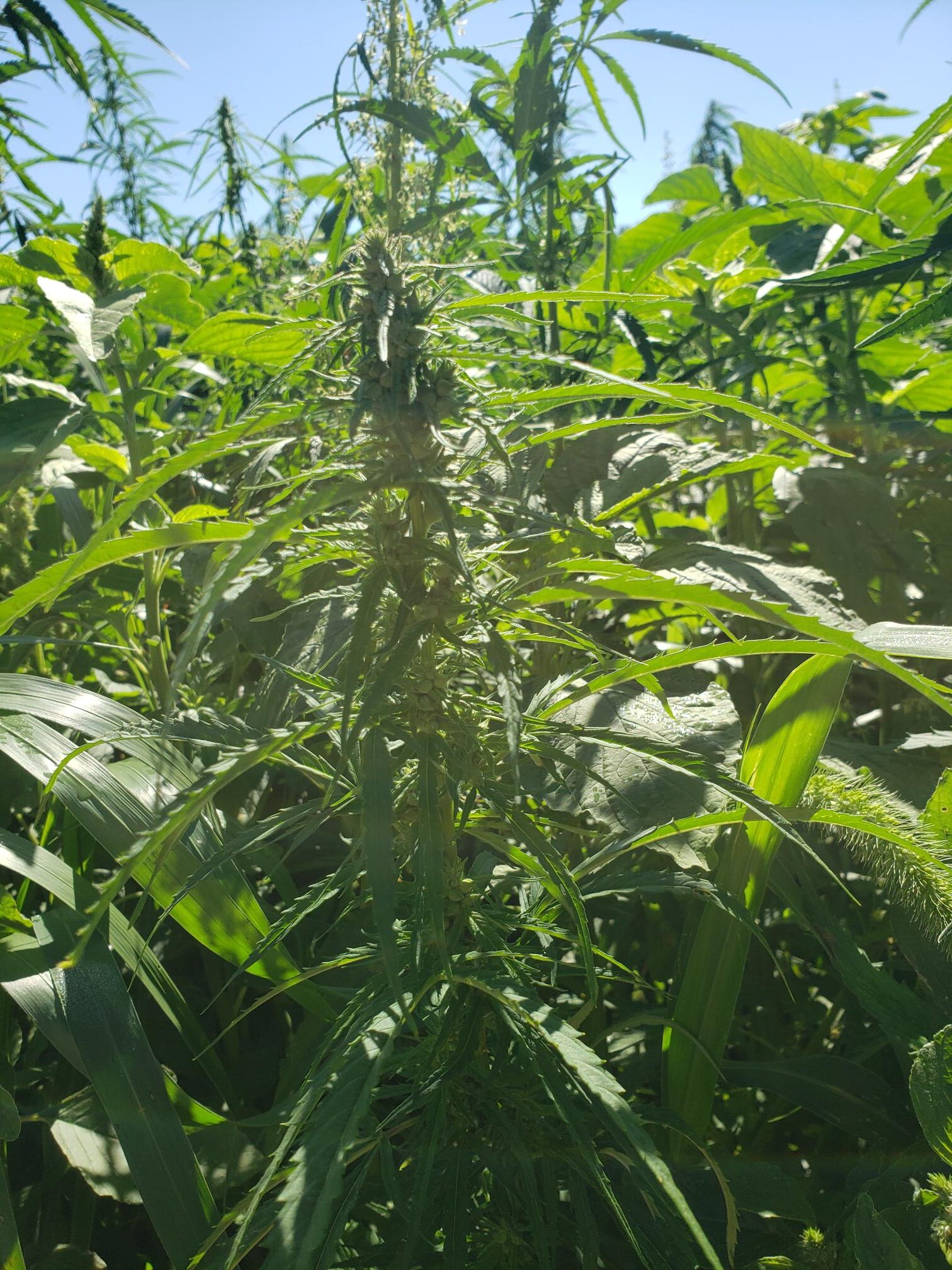
- Controlling invasive legume plants such as Kudzu (Pueraria montana) using biocontrol methods.
- Molecular characterization of pathogenic Fusarium species in soybeans and evaluation of the effectiveness of cover crops to control Fusarium disease incidence.
Here’s more about my research:
Mendis, H. C., Madzima, T. F., Queiroux, C. & Jones, K. M. (2016). Function of succinoglycan polysaccharide in Sinorhizobium meliloti host plant invasion depends on succinylation, not molecular weight. mBio, 7(3), e00606-16. DOI: 10.1128/ mBio.00606-16.
Mendis, H. C., Queiroux, C., Brewer, T. E., Davis, O. M., Washburn, B. K., & Jones, K.M. (2013). The Succinoglycan endoglycanase encoded by exoK is required for efficient symbiosis of Sinorhizobium meliloti 1021 with the host plants Medicago truncatula and Medicago sativa (Alfalfa). Molecular Plant-Microbe Interactions, 26(9), 1089-1105.
Mendis, H. C., Thomas, V. P., Schwientek, P., Salamzade, R., Chien, J. T., Waidyarathne, P., & De La Fuente, L. (2018). Strain-specific quantification of root colonization by plant growth promoting rhizobacteria Bacillus firmus I-1582 and Bacillus amyloliquefaciens QST713 in non-sterile soil and field conditions. PloS one. 13(2): e0193119. doi.org/ 10.1371/journal.pone.0193119.
I am working with students to examine questions related to the natural history, physiological ecology, and conservation of reptiles and amphibians. Specifically, I am interested in how these organisms interact with their environment and what factors may be negatively impacting populations. I work with students on local northern Missouri projects, including natural history, ecology, and chytrid fungus infection. I also work internationally examining insular dwarfism in Boa imperator in Cayos Cochinos, Honduras.
Check out these papers to learn more about research in the Montgomery lab:
Muelleman, P.J., O. DaCunha, and C.E. Montgomery. 2018. Crotalus horridus (Timber Rattlesnake) maternal scent trailing by neonates. Northeastern Naturalist 25: 50-55.
Taylor, H.L., A.J. Wilmes, C.E. Montgomery, L.J. Livo, and J.M. Walker. 2017. Recent northward range expansion of parthenogenetic Aspidoscelis tesselata in Colorado, a latitudinal baseline for the species, and a new hypothesis for the allopatry of pattern classes C and D at the northern periphery of the range. Southwestern Naturalist 62: 179-186.
Montgomery, C.E. 2017. Dwarfism in the Cayos Cochinos Boa Constrictor, Boa imperator. Serpens, 5: 4-5.
Check out these papers to learn more about research in the Patrick lab:
Erin I. Hayes and Joyce E. Patrick. yozG is needed for swarming motility in the undomesticated Bacillus subtilis strain NCIB 3610. Transactions of the Missouri Academy of Science. 49 (2022): 27–35.
Patrick, J.E. and D.B. Kearns. 2012. Swarming motility and the control of master regulators of flagellar biosynthesis. Molecular Microbiology. 83: 14-23.
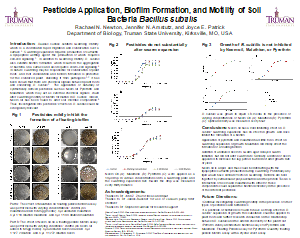
Newton, R., Amstutz, J. and Patrick, J.E. 2020. Biofilm formation by Bacillus subtilis is altered in the presence of pesticides. Access Microbiology.
Joyce E. Patrick and John Z. Zhu. Video Making Tips for Laboratory Instructors. CourseSource 9.
Dr. Michael Seipel
University of Missouri-Columbia
mseipel@truman.edu
Check out these papers to learn more about research in the Sieg lab:
Sieg RD, Hubbard JK, Penczykowski RM, Willard M, and Dwyer Z. 2023. A novel course-based experience to promote ecological field skills during the COVID-19 pandemic: Plantago as a tool for hybrid ecology coursework. Science Education and Civic Engagement: An International Journal. 15(1): 31-41.
Penczykowski RM and Sieg RD. 2021. Plantago spp. as models for studying the ecology and evolution of species interactions across environmental gradients. The American Naturalist. 198(1): 158-176.
Sieg RD, Wolfe K, Willey D, Kubanek J. 2013. Chemical defenses against grazers and fungi limit establishment of fungal farms on salt marsh angiosperms. Journal of Experimental Marine Biology and Ecology. 446:122-130.
Poulson K.L.*, R.D. Sieg*, E.K. Prince, J. Kubanek. 2010. Allelopathic compounds of a red tide dinoflagellate have species-specific and context-dependent impacts on phytoplankton. Marine Ecology Progress Series 416: 69-78. (* denotes joint first authorship).
Sieg R.D., D. Willey, K. Wolfe, J. Kubanek. 2013. Multiple chemical defenses produced by Spartina alterniflora deter farming snails and their fungal crop. Marine Ecology Progress Series 488: 35-49.
Check out these papers to learn more about research in the Weisstein lab:
Weisstein, A.E., E. Gracheva, Z. Goodwin, Z. Qi, W. Leung, C.D. Shaffer, and S.C.R. Elgin. 2016. A hands-on introduction to hidden Markov models. CourseSource 3.
Weisstein, A.E. 2011. Building mathematical models and biological insight in an introductory biology course. Mathematica Models of Natural Phenomenon 6: 198-214.
Weisstein, A.E. 2010. The case of the protective protein: Using a population genetics simulation in an undergraduate lab course to test hypotheses for the evolution of an HIV resistance allele. Biology International 47: 109-116.
Agricultural and Biological Sciences Department